Overview
Python-based data source templates use a fundamentally different approach to metadata ingestion than their SQL counterparts. Instead of separate SQL queries for each metadata level, a Python template uses a single script that connects to the data source programmatically and returns up to three pandas DataFrames — one each for schemas, tables, and columns.
This approach enables Validatar to discover metadata from sources that don't have a SQL interface: REST APIs, file systems, cloud storage, custom platforms, and anything else reachable through a Python library.
This article covers metadata ingestion for Script category templates. For SQL-based templates, see Metadata Ingestion Scripts — SQL Templates.
The Metadata Ingestion Tab — Python Templates
Open a Python-based template and navigate to the Metadata Ingestion tab. Unlike SQL templates, which show separate tabs for schema, table, and column levels, Python templates show a single script editor.
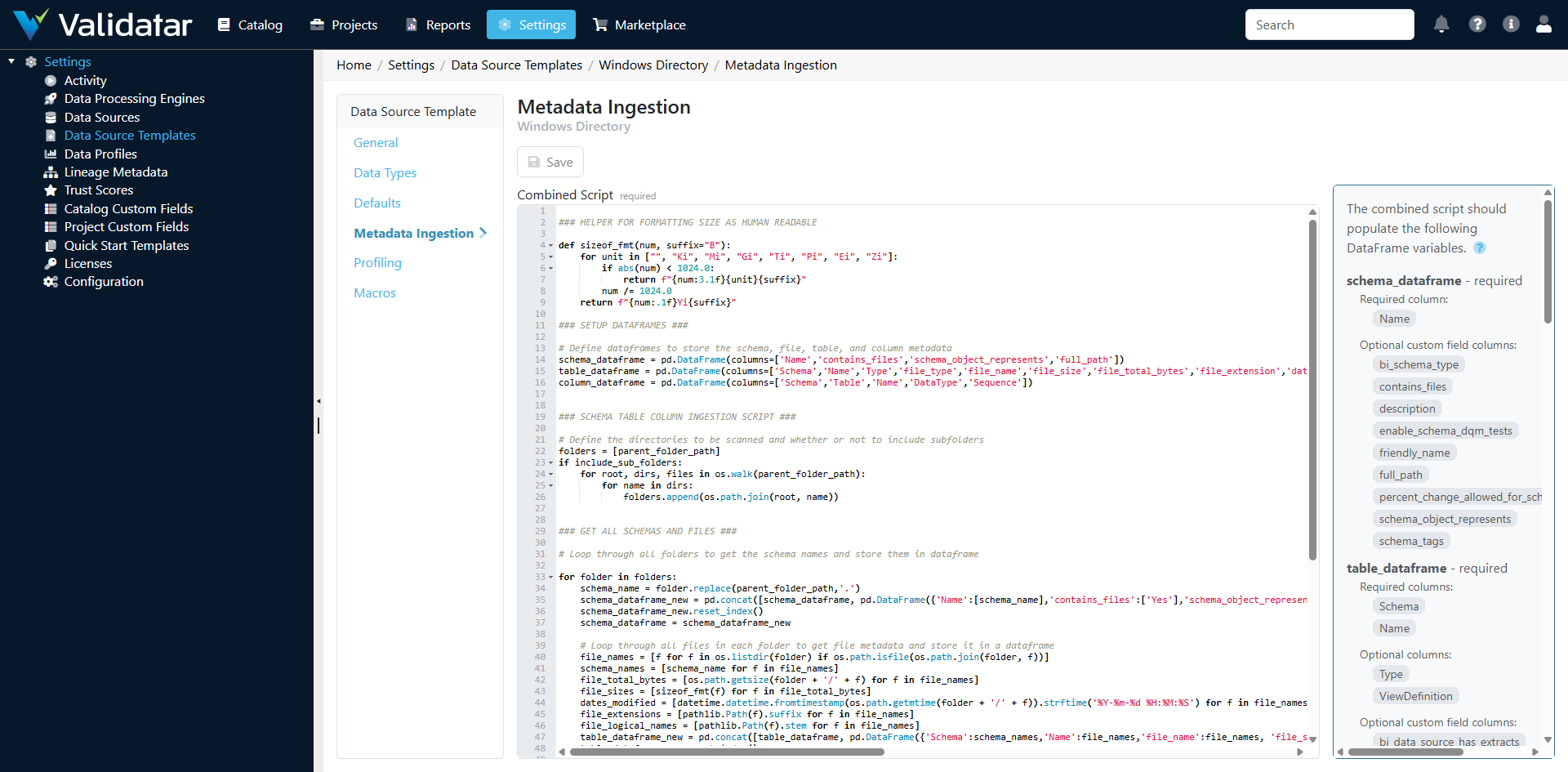
How Python Ingestion Works
The ingestion script is a Python script that Validatar executes through its embedded Python runtime. The script:
- Accesses template parameters (API keys, file paths, connection info) from the execution context
- Connects to the external data source using appropriate Python libraries
- Discovers the metadata structure
- Returns up to three pandas DataFrames with specific names and column structures
Validatar's Python runtime provides pandas, requests, and other common libraries. The script runs in a managed environment with access to the parameters defined on the template's Defaults tab.
Expected DataFrame Structure
The script should produce DataFrames with these specific names and columns. Not all three are required — return only the DataFrames relevant to your data source.
Schema DataFrame
DataFrame name: schemas
| Column | Required | Description |
|---|---|---|
schema_name |
Yes | The name of the schema or logical grouping |
Table DataFrame
DataFrame name: tables
| Column | Required | Description |
|---|---|---|
schema_name |
Yes | The schema this table belongs to |
table_name |
Yes | The name of the table, collection, or endpoint |
table_type |
No | Type classification (e.g., TABLE, VIEW, ENDPOINT) |
Column DataFrame
DataFrame name: columns
| Column | Required | Description |
|---|---|---|
schema_name |
Yes | The schema |
table_name |
Yes | The table |
column_name |
Yes | The column or field name |
data_type |
Yes | The data type (must match a mapping in the Data Types tab) |
ordinal_position |
No | Field order |
is_nullable |
No | Whether the field allows nulls (YES/NO) |
Accessing Parameters in Scripts
Template parameters defined on the Defaults tab are available in the script's execution context. This is how the script gets credentials, URLs, and other configuration values without hardcoding them.
# Parameters are available through the execution context
# The exact access pattern depends on the parameter reference keys
# defined on the template's Defaults tab
import pandas as pd
import requests
# Access parameters
base_url = parameters['api_base_url']
api_key = parameters['api_key']
Note: Secret parameters (like API keys) are decrypted at runtime and available in plain text within the script. The script executes in a secure, managed environment — but be mindful not to log or print sensitive values.
Example: REST API Ingestion Script
This example shows a complete ingestion script for a REST API that uses Swagger/OpenAPI to describe its endpoints:
import pandas as pd
import requests
import json
# Get parameters from template configuration
base_url = parameters['api_base_url']
api_key = parameters['api_key']
# Fetch the API schema (e.g., from a Swagger endpoint)
headers = {'Authorization': f'Bearer {api_key}'}
response = requests.get(f'{base_url}/swagger/v1/swagger.json', headers=headers)
spec = response.json()
# Build schema DataFrame
# For an API, "schemas" might represent API versions or resource groups
schema_records = []
for tag in spec.get('tags', []):
schema_records.append({'schema_name': tag['name']})
schemas = pd.DataFrame(schema_records)
# Build table DataFrame
# Each API endpoint becomes a "table"
table_records = []
for path, methods in spec.get('paths', {}).items():
for method, details in methods.items():
tags = details.get('tags', ['default'])
table_records.append({
'schema_name': tags[0],
'table_name': f'{method.upper()} {path}',
'table_type': 'ENDPOINT'
})
tables = pd.DataFrame(table_records)
# Build column DataFrame
# Response schema properties become "columns"
column_records = []
for path, methods in spec.get('paths', {}).items():
for method, details in methods.items():
tags = details.get('tags', ['default'])
# Extract response schema properties
responses = details.get('responses', {})
success_response = responses.get('200', responses.get('201', {}))
schema_ref = success_response.get('schema', {})
if 'properties' in schema_ref:
for i, (prop_name, prop_info) in enumerate(schema_ref['properties'].items()):
column_records.append({
'schema_name': tags[0],
'table_name': f'{method.upper()} {path}',
'column_name': prop_name,
'data_type': prop_info.get('type', 'string'),
'ordinal_position': i + 1,
'is_nullable': 'YES'
})
columns = pd.DataFrame(column_records)
Example: File System Ingestion Script
This example discovers metadata from a directory of CSV files:
import pandas as pd
import os
import csv
# Get parameters
directory_path = parameters['directory_path']
file_pattern = parameters.get('file_pattern', '*.csv')
# Schema = the root directory
schemas = pd.DataFrame([{'schema_name': os.path.basename(directory_path)}])
# Each file is a "table"
table_records = []
column_records = []
for filename in os.listdir(directory_path):
if filename.endswith('.csv'):
filepath = os.path.join(directory_path, filename)
table_name = os.path.splitext(filename)[0]
table_records.append({
'schema_name': os.path.basename(directory_path),
'table_name': table_name,
'table_type': 'FILE'
})
# Read header row to discover columns
with open(filepath, 'r') as f:
reader = csv.reader(f)
headers = next(reader)
for i, header in enumerate(headers):
column_records.append({
'schema_name': os.path.basename(directory_path),
'table_name': table_name,
'column_name': header,
'data_type': 'string', # CSV columns are string by default
'ordinal_position': i + 1,
'is_nullable': 'YES'
})
tables = pd.DataFrame(table_records)
columns = pd.DataFrame(column_records)
How Defaults and Overrides Work
The same inheritance model applies as with SQL templates:
- The template defines the default ingestion script
- Data sources using this template inherit the default
- Individual data sources can override the script with a custom version on their Schema Metadata page
Overrides are stored on the data source and don't affect the template or other data sources.
Tips for Writing Python Ingestion Scripts
Handle Errors Gracefully
Your script connects to external systems that may be unavailable, rate-limited, or returning unexpected responses. Use try/except blocks and return meaningful error messages:
try:
response = requests.get(url, headers=headers, timeout=30)
response.raise_for_status()
except requests.exceptions.RequestException as e:
raise Exception(f"Failed to connect to API: {str(e)}")
Respect Rate Limits
When ingesting from APIs, be mindful of rate limits. Add appropriate delays between requests if the API requires it. Metadata ingestion typically runs infrequently (daily or on-demand), so a few seconds of delay per request is acceptable.
Data Type Consistency
The data_type values in your column DataFrame must match the mappings on the template's Data Types tab. For API sources, map JSON types (string, integer, number, boolean, array, object) to corresponding entries in the Data Types configuration.
Return Empty DataFrames, Not None
If your data source doesn't have a concept of schemas (e.g., a single-endpoint API), still return a schemas DataFrame with at least one row. Use a logical grouping name like the API name or data source name.
Test Incrementally
When developing a new ingestion script:
- Start with just the schema DataFrame and verify it ingests correctly
- Add the table DataFrame and re-test
- Add the column DataFrame last
- Check the data catalog after each ingestion to verify the results
How This Fits Into the Bigger Picture
Python ingestion scripts are what make Validatar's "connect to any data, anywhere" promise a reality. While SQL templates cover traditional databases, Python templates extend that reach to APIs, files, cloud storage, and any other data source with a Python library.
Once metadata is ingested — regardless of whether it came from SQL or Python scripts — the downstream experience is identical: profiling, test recommendations, template test materialization, and macros all work the same way. The ingestion method is transparent to users working with the data source.
For Python-specific profiling configuration, see Profiling Configuration — Python Templates. For the complete end-to-end view, see Data Source Templates: The Complete Picture.